Logistic regression
What does logistic regression do?
- builts regression model when the dependent variable is categorical, with only two possible values (e.g. 0, 1)
- function:
glm()(Generalized Linear Models); by using the appropiate link function:
\[ \ln \left( \frac{p}{1-p} \right) = \beta_0 + \beta_1 \cdot x_1 + \beta_2 \cdot x_2 + \dots \]
\[ p = \frac{1}{1 + e^{ - \left( \beta_0 + \beta_1 \cdot x_1 + \beta_2 \cdot x_2 + \dots \right) }} \]
Assumptions
- underlying distribution is binomial
- plot should be S-shaped (always plot first!)
Logistic regression using glm()
Beetles data (for a given dosage of an insecticide: is beetle number x killed or not?):
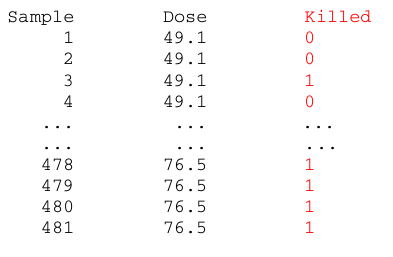
The data can be summarized by registering the portion of killed beetles:
beetles = read.table("beetles.txt", header = TRUE)
beetles## Dose Exposed Killed
## 1 49.1 59 6
## 2 53.0 60 13
## 3 56.9 62 18
## 4 60.8 56 28
## 5 64.8 63 52
## 6 68.7 59 53
## 7 72.6 62 61
## 8 76.5 60 60Add a column with the proportion of killed beetles:
Proportion = beetles$Killed / beetles$Exposed
beetles = cbind(beetles, Proportion)
beetles## Dose Exposed Killed Proportion
## 1 49.1 59 6 0.1016949
## 2 53.0 60 13 0.2166667
## 3 56.9 62 18 0.2903226
## 4 60.8 56 28 0.5000000
## 5 64.8 63 52 0.8253968
## 6 68.7 59 53 0.8983051
## 7 72.6 62 61 0.9838710
## 8 76.5 60 60 1.0000000plot(Proportion ~ Dose, data = beetles, main = "Proportion killed vs. dosage")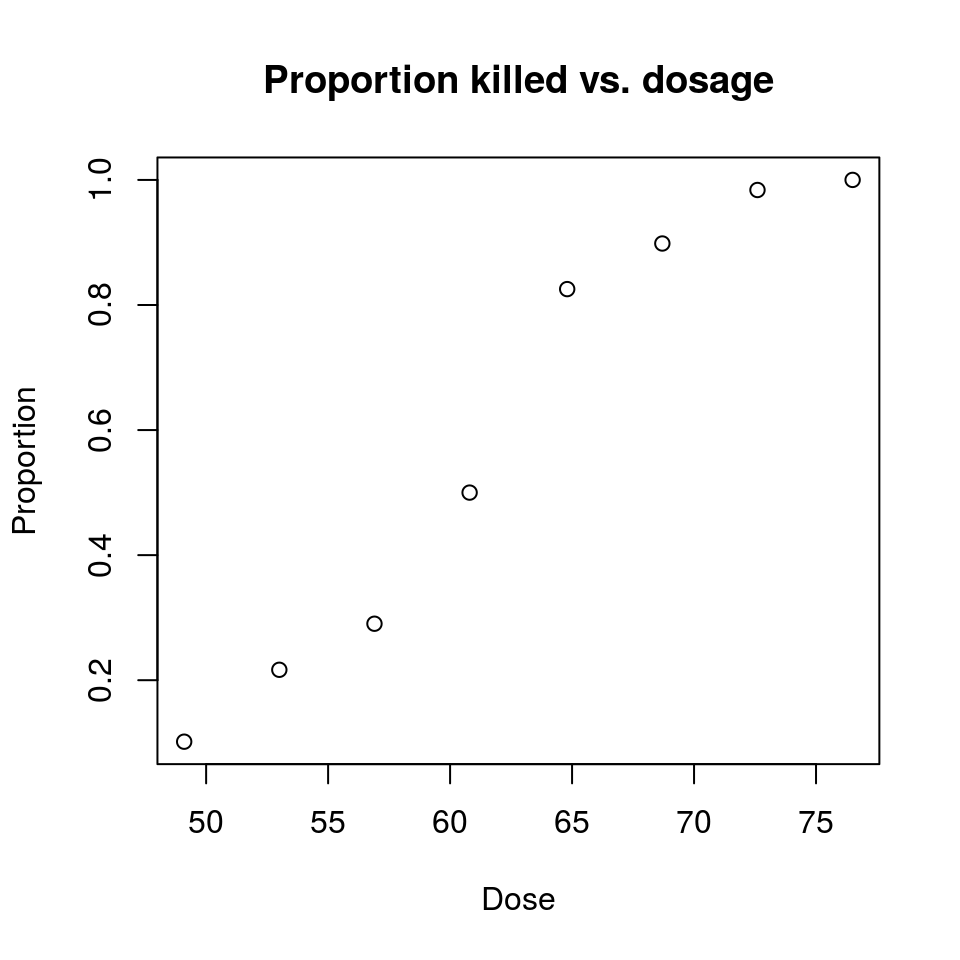
If this is an S-shaped function varying between 0 and 1, then logistic regression might work !
\(\begin{aligned} p = \frac{1}{1 + e^{ - t }} = \frac{e^t}{1 + e^t} \end{aligned}\)
Logistic function
- t can be any real number
- … but p is confined to (0,1)
survived = beetles$Exposed - beetles$Killed
killed = beetles$Killed
exposure = beetles$Dose
logfit = glm(cbind(killed, survived) ~ exposure, family = binomial) # logit link!
summary(logfit)##
## Call:
## glm(formula = cbind(killed, survived) ~ exposure, family = binomial)
##
## Deviance Residuals:
## Min 1Q Median 3Q Max
## -1.2746 -0.4668 0.7688 0.9544 1.2990
##
## Coefficients:
## Estimate Std. Error z value Pr(>|z|)
## (Intercept) -14.82300 1.28959 -11.49 <2e-16 ***
## exposure 0.24942 0.02139 11.66 <2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## (Dispersion parameter for binomial family taken to be 1)
##
## Null deviance: 284.2024 on 7 degrees of freedom
## Residual deviance: 7.3849 on 6 degrees of freedom
## AIC: 37.583
##
## Number of Fisher Scoring iterations: 4Reading the output
Coefficients / Estimate displays the calculated values for \(\beta_0\) and \(\beta_1\)
- \(\beta_0 = -14.82300\)
- \(\beta_1 = 0.24942\)
Coefficients / Pr(>|z|) are the p-values for the Wald-test
- if these p-values are low, the coefficients are significantly different from zero
- that means that there is a real dependency between the independent variables and the response
AIC: tells “how much information is lost” if the real data is replaced by the model ( Wiki)
Draw the regression line using the glm() results
beta.zero = coef(logfit)[1]
beta.zero # -14.82## (Intercept)
## -14.823beta.one = coef(logfit)[2]
beta.one # 0.25 ## exposure
## 0.2494156\[ p(x) = \frac{e^{-14.82 + 0.25x}}{1 + e^{-14.82 + 0.25x}} \]
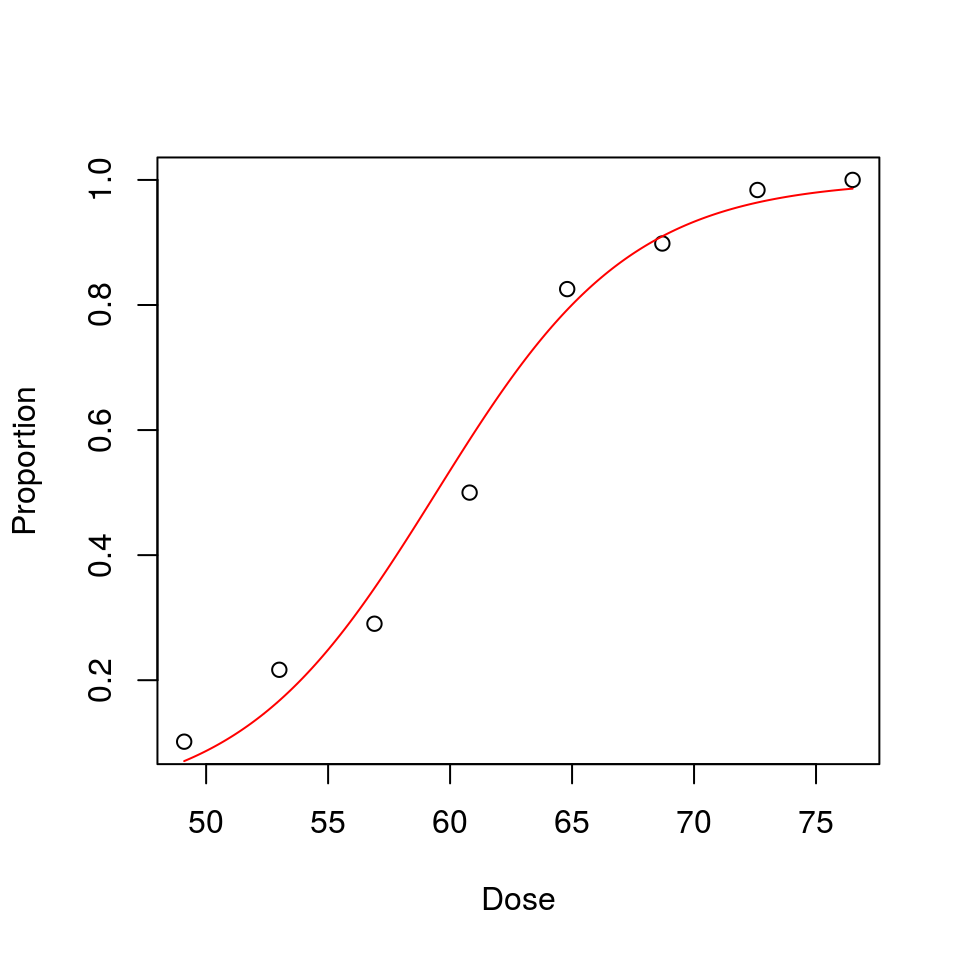
plot(Proportion ~ Dose, data = beetles) # observed
fitted <- function(x) { exp(beta.zero + beta.one*x)/(1 + exp(beta.zero + beta.one*x)) }
lowest.x = min(beetles$Dose)
highest.x = max(beetles$Dose)
curve(fitted(x), from = lowest.x, to = highest.x, n = 201, add = TRUE, col = "red")